How genetics and epigenetics link to periodontal disease: Research paper
By Felicia Hollander and Olive Njari
Biological sciences including genetics and epigenetics can be analyzed for how gene regulation affects the body's immune response to an inflammatory disease such as periodontitis. Genetics and epigenetics can be defined as the processes by which genes were expressed or suppressed through polymorphism, (de)methylation, or (de)acetylation.
Studies note that specific environmental factors such as race, gender, diabetes, education, smoking, and body mass index (BMI) all increase the severity of periodontal disease. One quarter of chronic periodontitis patients tested for hyper/hypo methylation proved to be a significant factor in response to inflammatory disease. The authors concluded while it was not genetics alone that affected response or susceptibility to periodontitis, it was environmental factors that also regulated gene transcription. These regulations stimulated cytokines such as IL-1 and IL-6, which were noted in inflammatory destruction of periodontal tissues (Loo, Jin, Cheung, Wang, & Chow, 2010).
This paper reports how controlling the enzymes that caused (de)methylation and (de)acetylation could be used as diagnostic tools and therapies that could help treat inflammatory diseases such as periodontitis (Gomez, Dutra, & Moreira, 2009).
Periodontitis
Periodontal diseases are bacterial infections that affect the tissues of the oral cavity. Periodontitis, a form of periodontal disease, destroys the tissues and the bone surrounding the teeth. Giannobile noted, “Periodontitis is a leading cause of tooth loss in adults, affecting more than 50 percent of the U.S. population” (Giannobile, 2012, pp 65). The Center for Disease Control and Prevention reported prevalence for periodontitis as “47.2% of adults aged 30 years and older have some form of periodontal disease. Periodontal disease increases with age; 70.1% of adults 65 years and older have periodontal disease” (CDC, 2015).
The numbers were significant in finding and led the authors to further research the linkage between factors such as genetics and epigenetics and susceptibility to periodontitis.
Genetics
Genetics is “the study of inheritance and inheritable traits as expressed in an organism’s genetic material” (Bauman, 2014). It was learned that children from the same parents had different traits which were inherited when genes that resided in their parents’ chromosomes underwent meiosis. Each child received 23 chromosomes from each parent.
If the order of the genes shifts during the meiosis process, this is termed as allelic variant. Those changes are very rare and not presented in many individuals (mutation). According to Kinane & Hurt, when allelic variant occurs in at least 1% of the population, it is termed genetic polymorphism and was considered to be a normal variant in the population (2003).
Polymorphism, the ability to exist in several different forms and variations in gene expression, provides the genetic basis for human variation. Each nucleotide in the DNA or RNA, is signified by single nucleotide polymorphisms (SNPs), which are considered mutual among individuals and is based on deletion, insertion, or exchange of a single nucleotide. For instance, nucleotide adenine (A) could have been replaced with the nucleotide Guanine (G) (Genetic Home Reference, 2015).
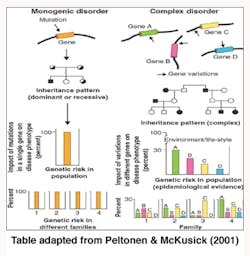
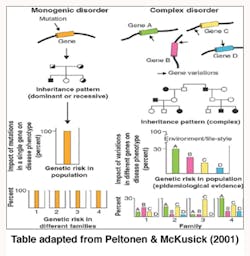
Genetics is termed as one of the causes of diseases and more so for the purpose of this report on periodontitis. The shifting of the nucleotides in the genes is said to influence periodontitis vulnerability. Some individuals are more susceptible to periodontitis than others (Tarannum & Faizuddin, 2012). “Geneticists have traditionally divided genetic diseases into two broad groups, Simple Mendelian diseases and complex diseases” (Kinane & Hart, 2003, pp. 431).
Alteration of a single gene locus causes Simple Mendelian diseases (monogenic disorders) and is known to follow the Mendelian mode of inheritance whereby one gene that was not normal was passed to an offspring by one parent (autosomal dominant) or two copies of a gene one from each parent that was abnormal was passed to the child (autosomal recessive) and caused diseases. Papillon Lefevre Syndrome (palmoplantar hyperkeratosis and aggressive periodontitis) was noted as an autosomal recessive disease and showed severe periodontal disease manifestations (Tarannum & Faizuddin, 2012).
Complex genetic diseases (polygenic disorders) are the result of altering more than one gene loci and are not known to follow the Mendelian mode of inheritance. Environmental factors such as aging, nutrition, and smoking play an important role in the complex genetic disease progression. Gene variations in polygenic disorders are considered to be genetic polymorphism as they work within the normal range of function (Kinane & Hart, 2003).
Periodontitis is considered a complex disease where a particular gene is not associated with the disease. It links to various genes, each contributing a small portion of the risk (Tarannum & Faizuddin, 2012).
Genetic Studies
Studies were conducted using different methods in the evaluation of genetic predisposition to periodontitis. Methods used were familial aggregation, twin studies, association studies, segregation, and linkage analysis. Familial aggregation studies focused on members of families who had a history of aggressive periodontitis. Familial studies conducted in Germany were reviewed by Hassell and Harris (1995) and concluded, “This aggregation within families strongly suggests a genetic predisposition” (Kinane & Hart, 2003, pp 435).
It was noted that other lifestyle and behavioral factors could have played part in the familial studies and may have influenced susceptibility to aggressive periodontitis. Chronic periodontitis, a less severe type of periodontal disease, is not evident until someone is 30 years of age, whereas the more severe form of periodontal disease, aggressive periodontitis, is evident earlier on in life. This difference in the onset signs of disease made diagnosis difficult, especially diagnosing adults’ type of aggressive periodontitis (Kinane & Hart, 2003).
A study of fraternal and identical twins living together or separately in Minnesota was conducted in 1991 by Hassell & O`Hehir. Age and gender were taken into account. Measurements of periodontal pockets, recession, and thickness of biofilm and severity of gingivitis were taken into consideration. It was concluded that “identical twins (whether reared together or apart) suggests that there is a genetic contribution to variations in levels of supragingival plaque and clinical measures of periodontal health” (Hassell & O`Hehir, nd). The fraternal twins whether living together or separately showed fewer similarities. In regard to the alveolar bone height measurements, environmental factors were shown to have influenced the results on both fraternal and identical twins living together or separately (Hassell & O`Hehir, nd).
Segregation and linkage analysis are noted as forms of scientific studies that look into the Mendelian mode of inheritance to determine if genetic traits of periodontitis exist. In 1994, Marazita and co-workers conducted a thorough segregation analysis on 100 North American families for aggressive periodontitis. It was concluded that “autosomal-dominant transmission with approximately 70% penetrance occurred for both Blacks and non-Blacks” (Tarannum & Faizuddin, 2012 pp 436).
Genetics and Susceptibility to Periodontitis
Vulnerability to periodontitis depends on the individual’s immune response. Defects of the tissues that line the body surface (epithelial tissue), tissues that connect, support and separate other tissues (connective tissues), and the cells that produce collagen in the connective tissues (fibroblasts) are noted to have contributed to the susceptibility of aggressive periodontitis. Decreased numbers of polymorphonuclear leukocytes (PMN) profoundly affect the host's susceptibility to periodontitis.
Cytokines noted as inflammatory responses influence the host response in periodontitis as well (Kinane & Hart, 2003). Cytokines are reported as powerful regulatory proteins released by the immune cells that influence the behavior of other cells (Nield-Gehrig & Willmann, 2011). Ianni, Bruzzesi, Pugliese, Porcellini, Carbone, Schiavone & Licastro, reported that different cytokines were involved in periodontitis. “SNPs of interleukin (IL-) 1α, IL-1ß, IL-4, IL-6, IL-8 and IL-18 located in different regions of the cytokine genes have been shown to affect the risk of the disease in several populations” (Ianni et al’s 2013, pp 2).
A chief emphasis is on the study of polymorphisms in the IL-1 due to the major part that IL-1β played in the development of periodontitis. Studies conducted suggested that individuals who had positive (higher levels) of IL-1 have a greater chance of having severe periodontitis than those who have negative (lower levels) of IL-1 (Tarannum & Faizuddin, 2012). “Periodontitis associated genotype (PAG) a composite of IL-1 genotype was found to be a combination of two rare alleles at separate SNPs at position -889 in the IL-1A promoter and at + 3954 (now referred to as +3953) of the IL-1B gene. Frequency of allele 2 of the IL-1B +3953 SNP proved significant increase in patients with advanced periodontitis” (Tarannum & Faizuddin, 2012, pp 12). Polymorphisms taken from genes of the IL-1 were tested for the relationship they had with generalized aggressive periodontitis (GAP). It was concluded that there was some evidence that GAP and the IL-1β polymorphisms are linked (Tarannum & Faizuddin, 2012).
More research was conducted to determine better ways of testing susceptibility to periodontitis much earlier on before an individual is infected by the disease. The test was performed on each patient who visited the dental office and who appeared to have healthy periodontal tissue. Blood (finger-stick) or saliva sample was collected and taken to the laboratory to be tested for PAG. If the results showed positive PAG it meant that the patient produced more IL-1 and was therefore susceptible to periodontitis. If the results showed negative PAG the patient produced less IL-1 and was less susceptible to periodontitis (Hassell & O`Hehir, nd).
Epigenetics
Many biologists from the nineteenth century believed that inheritance and development was one and the same. This theory was ignored for many years until it resurfaced and flourished in the twentieth century. Conrad Waddington, a professor of "Genetics at Edinburgh University" first coined the word epigenetics from the Greek word epigenesist. He was responsible for the birth of “Epigenetics Research Unit" at the university (Holliday, 2006, pp.76).
Following Waddington, many scientists discussed epigenetics; the material was reviewed by scientist Haig defining epigenetics as "observations that were not easily interpreted in genetic terms but had a heritable component, were liable to be labeled epigentic" (Holliday, 2006, pp. 77).
A more modern approach towards epigenetics was brought about by Yehuda & Bierer in 2009. During Yehuda & Bierer’s research it was discovered a link between trauma and the effects that it had on the offspring of those directly affected. Epigenetics was defined as environmental factors that altered gene expression in intergenerational terms. They studied the survivors of the Holocaust. They organized a center for counseling services similar to the Veterans Affairs (VA) for veterans. They discovered that it wasn't those directly involved with the Holocaust but their offspring that were coming forward and reaching out for counseling.
This led researchers to believe that there were deeper considerations of the trauma endured by these survivors. It was to their surprise that the survivor’s offspring had high levels of cortisol and consequently a difficult time dealing with stress-related issues. They attributed those findings to transgenerational effects that occurred from epigenetic (outside) factors that suppressed or expressed genes that regulated cortisol levels. These changes were noted as having been passed on to the Holocaust survivor’s offspring (Yehuda & Bierer, 2009). Similarly, a report described epigenetics as "changes in patterns of gene expression which do not involve changes in the DNA sequence” (Gomez, Dutra, & Moreira, 2009, pp. 625).
Modification of Histone Proteins and DNA Methylation
A gene is reported as the instructional portion of DNA that signals biological activity within our cells. DNA is also noted as being replicated through protein synthesis (transcription =>translation=>protein). DNA methylation is noted as chemical reactions that add a methyl group to a DNA molecule (hypermethylation) and demethylation (hypomethylation) as the removal of a methyl group from DNA (Yehuda & Bierer, 2009).
Histone proteins are responsible for packing and unpacking chromosomes for the transcription process. Histone modification results from enzymatic acetylation or deacetylation. These modifications are reported as chemical reactions that add or remove an acetyl group from the histone protein. Acetylation creates a less compact DNA complex that allows for gene expression. In contrast, deacetylation creates a more compact DNA complex that suppresses gene expression. It is said that there is a strong correlation between gene expression with both histone modification and DNA methylation (Wilson, 2008).
These types of reactions occur and cause alterations of gene expressions while changing the biological activity within the cells. It was important to note that this does not change the DNA. These type of expressions are site specific, reversible, and are termed epigenetic regulation (Yehuda & Bierer, 2009).
It was found that with methylation, demethylation and acetylation, deacetylation or any combination of these worked to either express or repress genes during replication. It was found that these processes were key factors in inflammatory and neoplastic diseases. These processes triggered by environmental influences regulated cytokines such as IL-1 and IL-6 which were responsible for inflammatory tissue destruction as seen in chronic periodontitis (Wilson, 2008).
Epigenetic Studies
Offspring receive both maternal and paternal copies of each gene during meiotic replication. On occasion (and what the research showed) there are times when only one of the genes is expressed. This is an example of epigenetic regulation. The most common is the X-chromosome, where only one is expressed in females. "The inactivated X-chromosome possesses high levels of DNA methylation, low levels of histone acetylation" are associated with gene suppression (Wilson, 2008, pp. 1516).
This suppression is seen within specific tissues, where inheritance came from one parent or the other, as well as differences within an organ itself. This appear as patches of certain cells expressed from the mother and in other areas of the same organ cells expressed from the father (Wilson, 2008).
The two biggest and most common epigenetic factors are nutrition and aging. Nutrition is reported as a factor that could be passed on from generation to generation through in utero exposure or through exposure from parental habits. Epigenetic effects from nutrition are seen in folic acid deficiency as well as deficiencies in selenium, arsenic, and polyphenols.
The diseases seen with these deficiencies were neural tube defects and cancers. While these play a significant part in gene expression they are reversible. Wilson reported on epigenetic factors with aging. The methylation and acetylation processes of 50 monozygotic twins were examined where very similar results were seen early on in life and substantial differences in later life. It was thought this was due to very similar lifestyles early on and significant differences as they aged (Wilson, 2008). Wilson reported in 2008 "that mounting evidence suggests that a disruption of the delicate balance in these epigenetic networks by environmental stimuli is a factor in a range of diseases, including cancer, inflammatory disorders, and autoimmunity"(p. 1516). "The Majority of studies have evaluated the methylation pattern in cancer, in which epigenetic silencing has been associated." Unfortunately, there are no studies directly linked to periodontitis (Gomez, Dutra, & Moreira, 2009, pp. 627).
Epigenetics and Susceptibility of Disease
The process of hyper or hypo methylation or histone acetylation, or any combination, brought forth the susceptibility of disease. Diseases of concern were inflammatory in nature, including breast cancer, rheumatoid arthritis, and periodontitis. In chronic periodontitis, hypermethylation of the genes E-Cardherin and Cyclooxygenase 2 cause an increase in susceptibility to inflammatory diseases including periodontitis.
A study from The Journal of Translational Medicine extracted DNA from the gingival tissues of 108 systemically healthy non-periodontitis subjects, blood samples of 110 chronic periodontitis patients, and neoplastic tissues of 106 breast cancer patients and tested for methylation of E-Cardherin and Cyclooxygenase 2 (Loo, Jin, Cheung, Wang, & Chow, 2010). The results were "hypermethylation of E-Cardherin and Cyclooxygenase 2 was observed in 38% and 35% of the breast cancer samples," "in Chronic Periodontitis patients the detection rate was 25% and 19%," "none was found in the systemically healthy non-periodontitis patients" (Loo, Jin, Cheung, Wang, & Chow, 2010, pp. 4), the findings were significant enough to associate hypermethylation to chronic periodontitis (Loo, Jin, Cheung, Wang, & Chow, 2010). In addition, Gomez, Dutra, & Moreira (2009) reported in preliminary studies that hypomethylation of IL-6 gene was found in people with periodontal disease.
The Center for Systemic and Oral Diseases reported that there were significant differences in response to therapy between patients with similar clinical exams. Biological aspects, immune response, and inflammatory response varied between patients. The biggest factors noted are environmental in nature and changed the phenotype and pathogenicity of each patient. Subject level variables that defined the levels of disease included race, gender, diabetes, education, smoking, and body mass index, all of which increased severity of periodontal disease. These variables interacted with a patient’s genetics and epigenetics, which influenced disease expression.
These factors are noted as contributors to the large range of immune response and inflammatory response at the cellular level. Epigenetics are critical regulatory factors in inflammatory response. Epigenetics are reversible and determined via environmental exposure, which may be inherited due to in utero exposure.
Epigenetic changes were passed from parental chromosomes in the gamete or zygote, thus altering gene expression of the somatic cells. These changes were through family history or environmental exposure. These effects were in all cells or in specific tissues within the body. These were considered transmissible since the change in expression occurs following cell division (Offenbacher, Barros, & Beck, 2008).
Implementation
Patients with genetic factors that increased cytokine levels such as IL-1 marked as higher risk for periodontitis (Tarannum & Faizuddin, 2012). Giannobile discussed a salivary test that tested for cytokines (interleukin) known to be associated with inflammatory disease. The test was reported as being available at chairside and would allow hygienists to recognize those with increased risk to periodontitis.
The dental practice needs to include salivary testing as part of routine dental diagnosis. This would allow for early identification of at risk patients, and allow for individualized care for those at a higher risk for periodontal disease (2012). The authors believe that with a thorough health history and nutritional assessments hygienists can identify environmental factors that increase a patient’s risk to inflammatory diseases. By altering or eliminating these epigenetic factors, the authors’ thoughts are that it would in turn change a person’s susceptibility to inflammatory diseases including periodontitis.
Conclusion
Vulnerability to periodontitis is increased when IL-1 gene polymorphisms occur and increase the levels of IL-1. It is hoped by the authors that through the introduction of chairside salivary testing, periodontitis susceptibility could be reduced and a personalized care plan could be implemented.
Epigenetics, an environmental exposure, chemically changed the expression of DNA transcription. Through these epigenetic factors—primarily diet and aging followed by smoking, diabetes, gender, BMI, race and education—an individual's predisposition for immune or inflammatory response is determined. It was learned that through (de)methylation or (de)acetylation such changes occurred.
These factors (if experienced while conceiving or in utero) alter the same genes of one's offspring, thus making an epigenetic change in DNA heritable or transmissible (Yehuda &Bierer, 2009). Epigenetic factors show increased severity of periodontitis, an inflammatory disease over those who were absent of these factors (Wilson, 2008). In conclusion, it is hoped that through epigenetic manipulation that controls the enzymes that increases production of cytokines, including IL-1 and IL-6, that it would become therapeutic for many inflammatory diseases, including periodontitis (Gomez, Dutra, & Moreira, 2009).
References
- Bauman RW. (2014). Microbiology with diseases by taxonomy (4th ed., p. 194). Glenview, IL: Pearson
- Egger G, Liang G, Aparicio A, Jones PA. (2004). Epigenetics in human disease and prospects for epigenetic therapy. Nature, 429(6990), 457-463.
- Giannobile WV, (2012), Salivary diagnostics for periodontal diseases The Journal of the American Dental Association, Volume 143, pp. 6S-11S doi: 10.14219/jada.archive.2012.0341
- Gomez RS, Dutra WO, Moreira PR. (2009). Epigenetics and periodontal disease: Future perspectives. Inflammation Research, 58(10), 625-629. doi:10.1007/s00011-009-0041-7
- Hassell T, O`Hehir T. Genetics & Periodontal Disease. RDH Magazine, 20(1). Retrieved November 2, 2015, from http://www.rdhmag.com/articles/print/volume-20/issue-1/feature/genetics
- Holliday R. (2006). Epigenetics: A historical overview. Epigenetics, 1(2), 76-80.doi/pdf/10.4161/epi.1.2.2762
- Ianni M, Bruzzesi G, Pugliese D, Porcellini E, Carbone I, Schiavone A, Licastro F. (2013). Variations in inflammatory genes are associated with periodontitis. Immunity & Ageing: I & A, 10, 39. doi.org/10.1186/1742-4933-10-39
- Kinane DF, Hart TC. (2003). Genes and gene polymorphisms associated with periodontal disease. Critical Review Oral Biology Medicine, 14(6), 430-449. doi:10.1177/154411130301400605
- Loo WT, Jin L, Cheung MN, Wang M, Chow LW. (2010). Epigenetic change in E-cadherin and COX-2 to predict chronic periodontitis. Journal of Translational Medicine, 8, 110. doi:10.1186/1479-5876-8-110
- Nield-Gehrig JS, Willmann DE. (2011). Foundations of periodontics for the dental hygienist (3rd ed., pp. 157-169). Philadelphia, PA: Wolters Kluwer Health.
- National Institutes of Health, What are single nucleotide polymorphisms (SNPs)? (October 26, 2015). In Genetic Home Reference. Retrieved November 1, 2015, from http://ghr.nlm.nih.gov/handbook/genomicresearch/snp
- Offenbacher S, Barros SP, Beck JD. (2008). Rethinking periodontal inflammation. Journal of Periodontology, 79(8S), 1577-1584. doi:10.1902/jop.2008.080220
- Periodontal Disease (2015, March 10). In Centers for Disease Controls and Prevention. Retrieved November 21, 2015 from http://www.cdc.gov/oralhealth/periodontal_disease/
- Pourabbas R, Kashefimehr A, Rahmanpour N, Babaloo Z, Kishen A, Tenenbaum HC, Cheng YSL. (2005). Position paper: Epidemiology of periodontal diseases. Journal of Periodontology, 76(8), 1406-1419. doi:10.1902/jop.2005.76.8.1406
- Taba M, Scombatti de Souza SL, Mariguela VC, (2012). Periodontal disease: a genetic perspective. Brazilian Oral Research, 26(spe1), 32-38. doi: 10.1590/S1806-83242012000700006
- Tarannum F, Faizuddin M. (2012). Effect of gene polymorphisms on periodontal diseases. Indian Journal of Human Genetics, 18(1), 9–19. http://doi.org/10.4103/0971-6866.96638
- Wilson AG. (2008). Epigenetic regulation of gene expression in the inflammatory response and relevance to common diseases. Journal of Periodontology, 79(8S), 1514-1519. doi:10.1902/jop.2008.080172
- Yehuda R, Bierer LM. (2009). The relevance of epigenetics to PTSD: Implications for the DSM-V. Journal of Traumatic Stress, 22(5), 427–434. doi:10.1002/jts.20448